Pseudotyping Bacteriophage P2 Tail Fibers to Extend the Host Range for Biomedical Applications | ACS Synthetic Biology
Receptor Diversity and Host Interaction of Bacteriophages Infecting Salmonella enterica Serovar Typhimurium | PLOS ONE

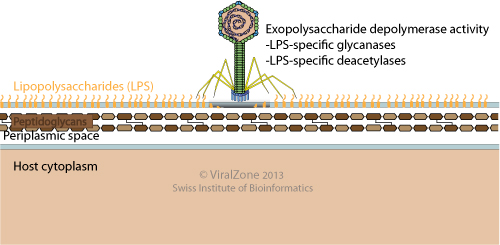
Novel Host Recognition Mechanism of the K1 Capsule-Specific Phage of Escherichia coli: Capsular Polysaccharide as the First Receptor and Lipopolysaccharide as the Secondary Receptor | Journal of Virology
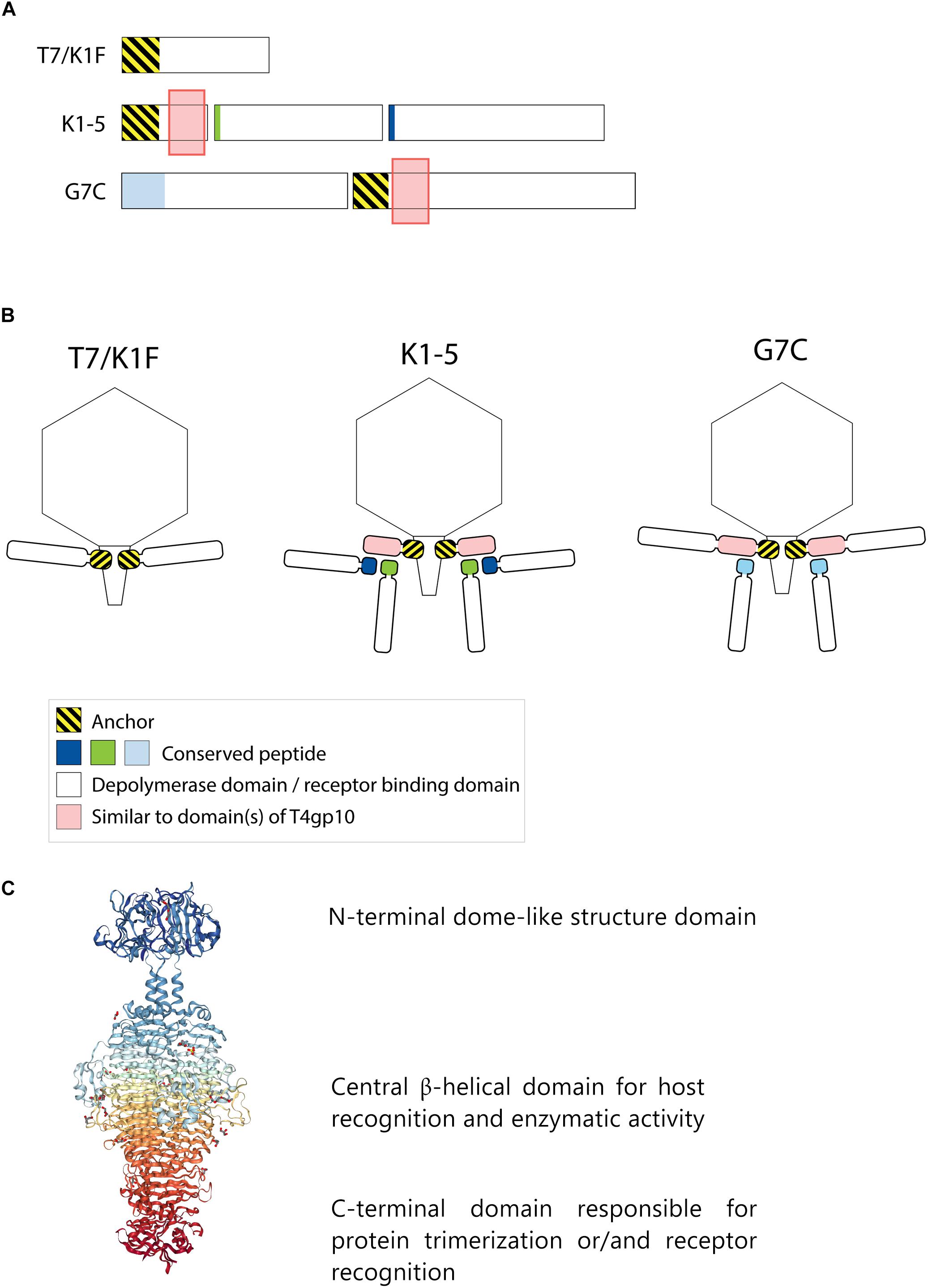
Frontiers | Modeling the Architecture of Depolymerase-Containing Receptor Binding Proteins in Klebsiella Phages

Phage–antibiotic combinations: a promising approach to constrain resistance evolution in bacteria - North - 2021 - Annals of the New York Academy of Sciences - Wiley Online Library

Pseudotyping Bacteriophage P2 Tail Fibers to Extend the Host Range for Biomedical Applications | ACS Synthetic Biology

Exploitation of a Klebsiella Bacteriophage Receptor-Binding Protein as a Superior Biorecognition Molecule | ACS Infectious Diseases
Molecular anatomy of the receptor binding module of a bacteriophage long tail fiber | PLOS Pathogens

A continuous evolution system for contracting the host range of bacteriophage T7 | Scientific Reports

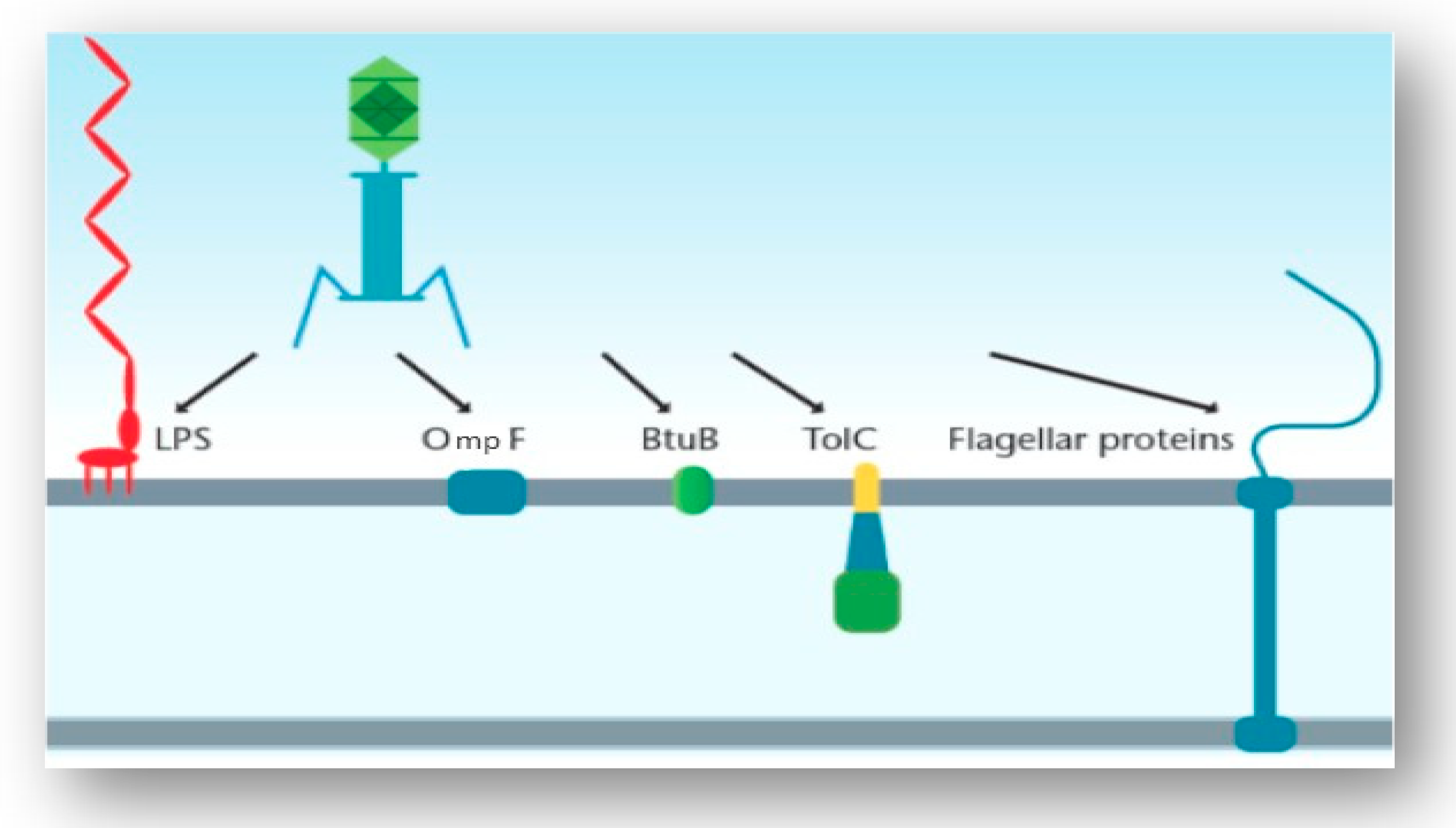
Receptors of Salmonella phages. Phages can use a number of cell surface... | Download Scientific Diagram

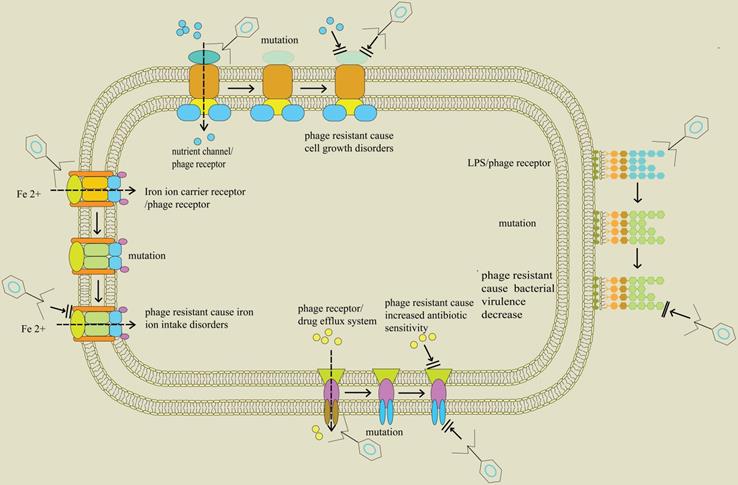
Examples of the possible host cell receptors a tailed phage generally... | Download Scientific Diagram
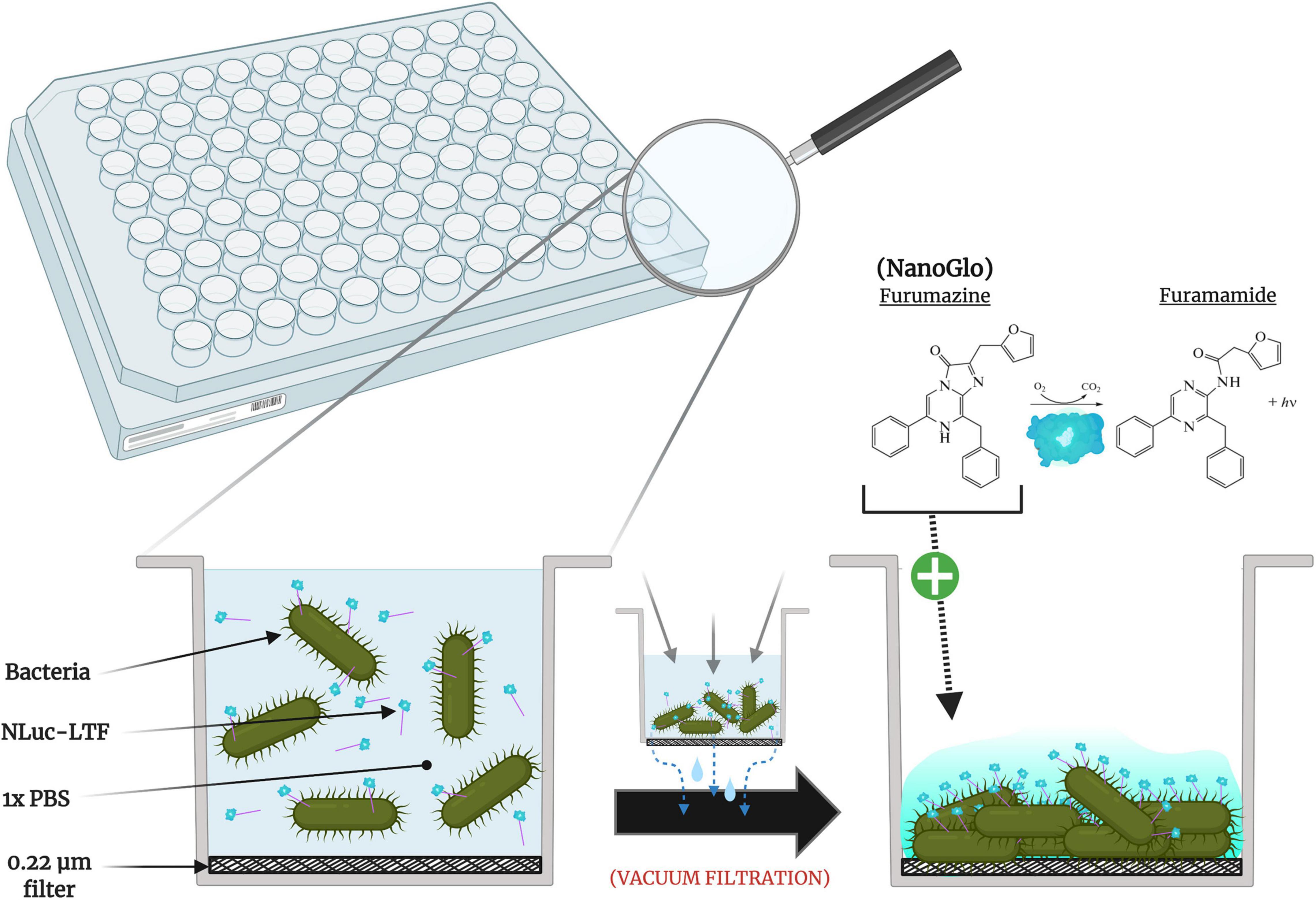
Frontiers | Evaluating Phage Tail Fiber Receptor-Binding Proteins Using a Luminescent Flow-Through 96-Well Plate Assay

Characterization of the interactions between Escherichia coli receptors, LPS and OmpC, and bacteriophage T4 long tail fibers - Washizaki - 2016 - MicrobiologyOpen - Wiley Online Library

High-throughput discovery of phage receptors using transposon insertion sequencing of bacteria | PNAS

Interactions of bacteriophage T4 adhesin with selected lipopolysaccharides studied using atomic force microscopy | Scientific Reports

Novel Host Recognition Mechanism of the K1 Capsule-Specific Phage of Escherichia coli: Capsular Polysaccharide as the First Receptor and Lipopolysaccharide as the Secondary Receptor | Journal of Virology

Systematic Discovery of Salmonella Phage-Host Interactions via High-Throughput Genome-Wide Screens | bioRxiv

A lipopolysaccharide-dependent phage infects a pseudomonad phytopathogen and can evolve to evade phage resistance | bioRxiv

The Presence of Two Receptor-Binding Proteins Contributes to the Wide Host Range of Staphylococcal Twort-Like Phages | Applied and Environmental Microbiology
Fitness Trade-Offs Resulting from Bacteriophage Resistance Potentiate Synergistic Antibacterial Strategies